INTRODUCTION
Shiga toxin (Stx)-producing Escherichia coli (STEC) cause gastrointestinal diseases such as Hemorrhagic colitis (HC), and in some cases this develops into the life-threatening hemolytic uremic syndrome (HUS).1,2 Foodborne pathogen, E. coli O157:H7, is a genetically heterogeneous serotype of STEC. Consumption of as low as ten cells of O157:H7 derived from contaminated bovine origin foods such as beef and cheese, people may develop symptoms ranging from asymptomatic carriage, bloody diarrhea, to fatal HUS. It has been estimated to cause 73,000 illnesses and 60 deaths annually in United States (US), with an estimated economic loss of about 400 million dollars. This pathogen continues to cause most of known outbreaks across the globe.3,4 Therefore, the US Department of Agriculture (USDA) has enforced a “zero tolerance policy” to regulate the prevalence of O157:H7 in foods. Genetic methods, most commonly the lineage specific polymorphism assay (LSPA)5 and clade typing,6 have been used to separate O157:H7 strains by virulence potential and ecology. For example, human isolates are more commonly classified as lineage I and I/II than lineage II,7,8 and it has been argued that isolates from clades 6 and 8 are more virulent than isolates from other clades.6,9,10
Many virulence factors contribute to the pathogenesis of E. coli O157:H7 including Stx.11,12 This is an AB5 toxin, comprised of a single A subunit associated with 5 identical B subunits.13,14 While the pentamer binds to Gb3 located in the host cell membrane,15 the A subunit functions as a glycosidase, cleaving an adenine nucleotide from 28S rRNA within the 60S subunit. This results in inhibition of protein synthesis, followed by necrosis and cell death.16 There are 2 immunologically distinct isoforms of the toxin, designated Stx1 and Stx2, which share 56.8% amino acid identity.17 Stx2 has a lower affinity for Gb3 than Stx118; however, Stx2 is more toxic in animal models19,20,21 and associated with more severe clinical cases.22 Among the seven subtypes of Stx2 (from Stx2a to Stx2g),23 Stx2a and Stx2c are more often associated with clinical strains causing deadly HUS.24,25 In this manuscript, we will use “Stx2” when collectively referring to its subtypes.
Stx2 producing strains are correlated with the development of HUS,26,27 leading to the hypothesis that high toxin-producing strains are more likely to cause severe disease symptoms. Several studies have concluded, for example, that lineage I and I/II isolates produce more Stx2 than lineage II isolates,28 and that clade 8 isolates are high toxin producers compared to isolates from other clades.29,30 Most studies measure Stx2 production by semi-quantitative commercial kits,10 qPCR/microarray,10,29,30 or by semi-quantitative western blots.30 Concerning mRNA quantification, several studies have reported that Stx2 transcript levels do not always correlate with Stx2 toxin production.10,30,31 Thus, immunoassays are preferred methods for quantifying Stx production.
Several immunologic methods were previously described that use either immobilized antibodies specific for Stx,32,33 or a receptor mimic34,35,36 as the antigen capture in the sandwich ELISA. The disadvantage for the former is that it requires two sets of anti-Stx2 antibody from different species for both antigen capture and detection steps, while the latter assay only needs one set of anti-Stx2 antibody for detection. Most commercially available anti-Stx2 antibodies are monoclonal and generated from mice, so we suggest that one drawback preventing most research laboratories from quantifying Stx2 by sandwich ELISA is the limited commercially availability of antibodies from other species. Although, commercial kits such as Premier® Enterohemorrhagic Escherichia coli (EHEC) has been developed for detecting Stx, they are not designed to be specifically for Stx2 quantification, because they can not differentiate Stx1 from Stx2. Its high cost also prohibits most laboratories from routinely using it for this purpose. Moreover, previously reported sandwich ELISA by using Gb3 as antigen capture was not developed to be quantitative for Stx2. Therefore, we were motivated to develop a cheap and reliable assay specifically for quantifying Stx2a that required only one set of anti-Stx2a antibody. Our initial attempts to develop a R-ELISA met with several difficulties in consistency and repeatability, an issue echoed in a previous publication.36 We suggest here that development of a robust assay would require identifying and standardizing critical steps of this assay, so it can be successfully applied to quantify Stx2a.
MATERIALS AND METHODS
Strains and Culture Conditions
The fourteen E. coli O157:H7 strains used in this study were obtained from the Pennsylvania Department of Health and previously characterized by our lab.7 They were stored in 10% glycerol at -80 °C.
Ciprofloxacin Induction and Bacteria Lysate Collection
Each strain was propagated in Luria Broth (LB) at 37 °C with shaking overnight. Cells were diluted to A600 of 0.05 in fresh LB broth. To induce toxin expression, ciprofloxacin was added to a sub-leathal concentration of 45 ng/mL. After an eight-hour incubation at 37 °C with shaking, the culture was centrifuged at 4,000 g for 10 min, and supernatants were filtered through 0.2 µM cellulose acetate filters (VWR, Radnor, PA, USA).
Stx2 Specific R-ELISA
A 0.5 mg/ml stock solution of hydroxyl acyl ceramide trihexoside (Matreya Biosciences, Pleasant Gap, PA, USA) was prepared in chloroform: methanol (2:1, v/v). This stock solution was further diluted to 25 µg/mL in methanol, and was either used immediately or stored at -20 °C. Frozen stocks were heated in a 55 °C water bath to re-dissolve hydroxyl acyl CTH before use. To coat wells with antigen capture, a volume of 100 µL hydroxyl acyl CTH working solution was added to wells of an eight-well polystyrene strip plate (Thermo Scientific, Waltham, MA, USA) in a chemical fume hood. Strip plates were rotated manually every 15 minutes until the methanol was fully evaporated. Next, 200 µL blocking buffer [4% bovine serum albumin (BSA) in 0.01 M phosphate buffer saline (PBS) with 0.05% Tween20] was added to each well and incubated at 4 °C for 16 hours. After removing blocking buffer, 200 µL washing buffer (0.01M PBS with 0.05% Tween20) was added to each well and incubated for one minute on a shaking platform. This step was repeated for a total of 5 washes.
The R-ELISA assay was initiated by adding 100 µL sample into each well, followed by shaking at room temperature for one hour. Wells were washed five times in PBS/Tween20 as described above. Next, 100 µL mouse anti-Stx2 antibody (Santa Cruz Biotech, Santa Cruz, CA, USA) was diluted in blocking buffer to 1 µg/mL, added to each well, and incubated with shaking at room temperature for one hour. Wells were again washed five times with PBS/Tween20. Goat anti-mouse HRP conjugated secondary antibody was diluted in the blocking buffer (1 μg/mL, 100 µL) and added to each well, and incubated with shaking at room temperature for one hour. Five washes in PBS/Tween20 followed; however, this time each wash was performed for five minutes. Detection was accomplished using 1-Step Ultra TMB (Thermo-Fischer, Waltham, MA, USA), which was equilibrated to room temperature in a foil-wrapped tube for at least 30 minutes prior to use. TMB substrate (100 µL) was added to each well and incubated with shaking for 10 minutes to allow for color development. Finally, 100 µL of stop solution (2 M H2 SO4 ) was added to each well, followed by shaking for 30 seconds. The reading values of A450 were obtained using a DU®730 spectrophotometer (Beckman Coulter, Atlanta, Georgia, USA).
For all assays, supernatants from E. coli O157:H7 strain PA24, which produces only Stx1, were used for the negative control, and a lysate from a E. coli O157:H7 strain, PA11, served as the positive control. The Stx2a concentration for PA11 stock was quantified by known concentration of pure Stx2 (BEI Resources, Manassas, VA, USA). The standard curves in R-ELISA were generated using two-fold serially diluted PA11 lysate in PBS. Any A450 above 0.1 was considered as positive. Total protein in each unknown sample was measured by the Bradford assay (VMR Life Science, Philadelphia, PA, USA), following the manufacturer’s recommended protocol. Stx2 quantities were reported as µg Stx2/mg total protein.
Statistic Analysis
All data were reported as the mean from three biological replicates and each sample had three technical repeats. The mean and standard error were calculated in MS Excel®. Data were analyzed using linear regression and one-way analysis of variance (ANOVA) in Minitab version 17 (Minitab Inc., State College, PA, USA).
RESULTS
Assay Optimization
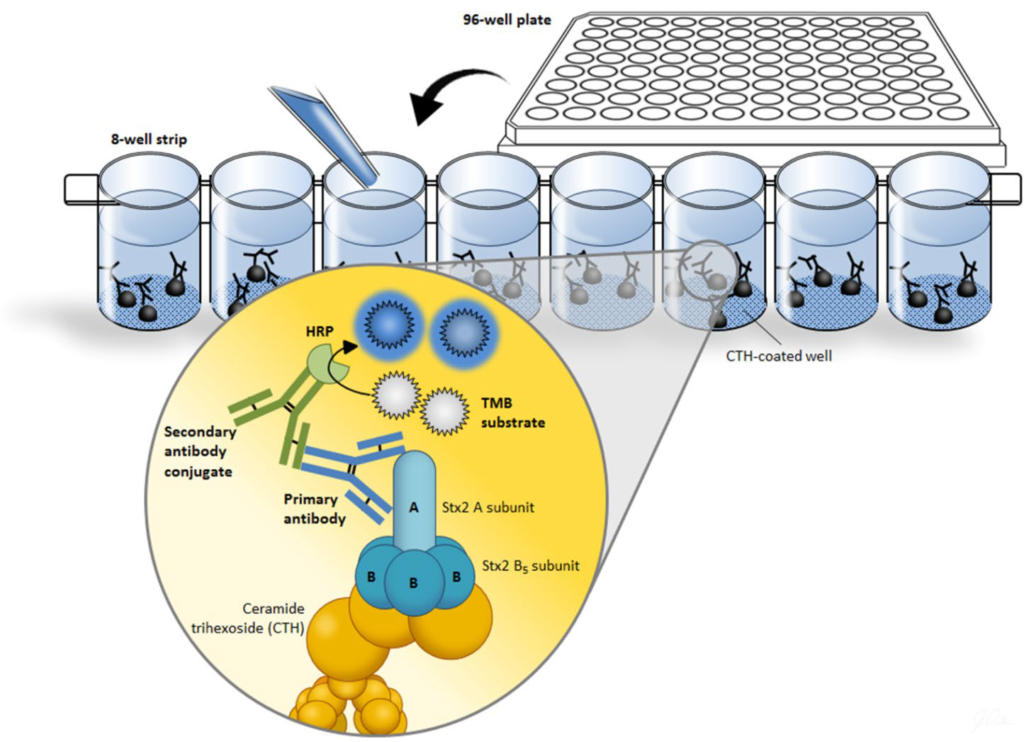
We started to develop our R-ELISA based on the traditional sandwich ELISA. Although, polyclonal anti-Stx2 antibody from rabbit (BEI resource, Manassas, VA, USA) was tried as the antigen capture in our initial attempt, it gave very high background noise. Therefore, we replaced the capture antibody with a Gb3 chemical analogue, hydroxyl acyl ceramide trihexoside (CTH) (Figure 1). Mouse anti-Stx2 A subunit monoclonal antibody (Santa Cruz Biotech, Santa Cruz, CA, USA) was selected as the primary antibody. As a result, we were able to reduce the background noise, by giving optical density at 450 nm (A450) less than 0.1 when measuring the negative control.
Figure 1: Schematic of R-ELISA used to Detect Stx2. The Stx2 Capture Hydroxyl acyl CTH is Added to the Detachable Eight-Well Polystyrene Strip Plate, to which Antigen Stx2 Specifically Binds. Primary Antibody Binds Specifically to Stx2. A Horseradish Peroxidase (HRP)-Conjugated Secondary Antibody is Added, and a Positive Reaction is Detected by a Color Change after Addition of 3,3’,5,5’ Tetra Methyl Benzidine (TMB).
Using previous publications34,36 as guidelines, several critical steps which were not explicitly stated in those manuscripts were identified and further improved to make our assay robust and quantitative. First, although several Gb3 receptor mimics are available (CTH is available as hydroxyl acyl, nonhydroxyl acyl and lyso-derivatives), we found that hydroxyl acyl CTH was best at capturing Stx2a at room temperature (Table S1). Second, although the supplier recommended methanol as the solvent, we found that dissolving hydroxyl acyl CTH first in chloroform:methanol (2:1), and further diluting in methanol to the working concentration gave the best results. Commercial hydroxyl acyl CTH better dissolved in the former without leaving any solid residual, and the signal got improved by average five times higher (Table S2). Third, we used detachable eight-well polystyrene strip plates instead of traditional 96-well plates. This minimized edge effects, a common observation with ELISA where peripheral wells give a higher absorbance than central wells. Fourth, the coating of R-ELISA plate wells with hydroxyl acyl CTH needs to be performed as rapidly as possible, so evaporation of methanol occurs evenly across the wells. Last, each of the washing steps used to remove the secondary antibody was extended to five minutes to minimize background noise.
Specificity and Sensitivity of Stx2 Specific R-ELISA
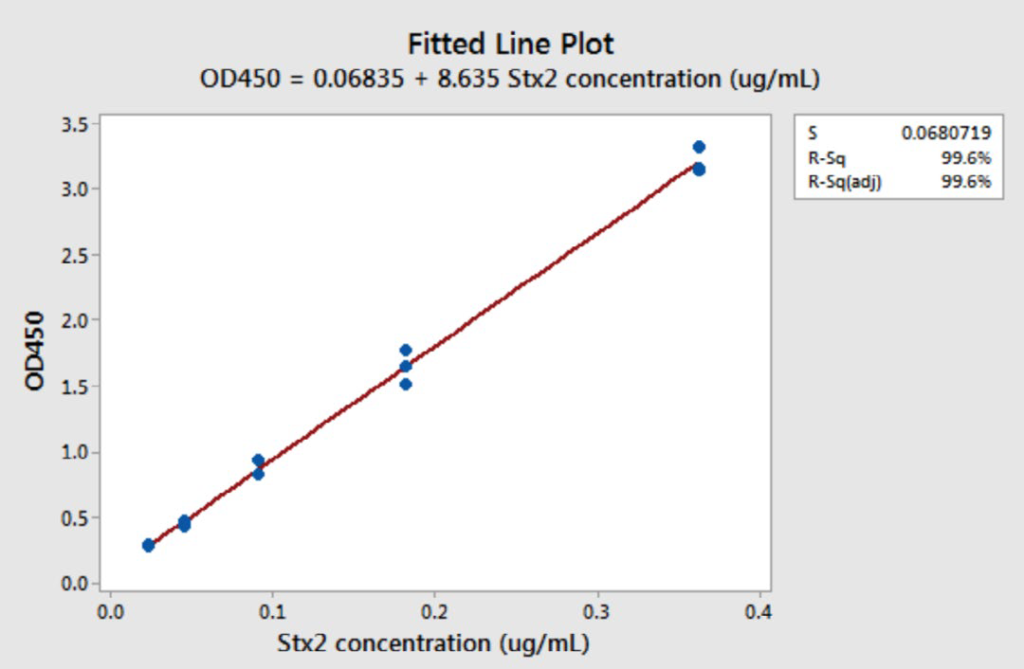
The specificity of this assay was first tested by using bacteria lysate of Stx1-producing strain previously designated PA24.7 The absorbance at 450 nm (A450) for bacteria lysate from PA24 is routinely less than 0.1, indicating that no cross-reactivity was observed in this assay with Stx1. As pure Stx2a from biodefense and emerging infectious (BEI) resource is available in quite limited quantity, we decided to use lysates from a high Stx2aproducing strain previously designated PA117 as the toxin source for standard curves. The Stx2a concentration in PA11 supernatants after ciprofloxacin induction was first quantified by using known concentration of pure Stx2a and around 23.2 µg/mL. We routinely obtained a linear range between an A450 of 0.3 to 3.2, corresponding to a Stx2a concentration from 23 ng/mL to 363 ng/mL (Figure 2).
Figure 2: Linear Curves Generated Using Bacteria Lysate Supernatant from E. coli O157:H7 Strain PA11 in R-ELISA. Two-fold Dilutions were Assayed using R-ELISA, Revealing the Linear Ranges of the Assay. The Linear Equation of the Best Fit Line and R-square are Shown by using Raw Data from Three Replicates.
Application of Stx2 Specific R-ELISA to Quantify Toxin in Bacteria Lysates
This assay was designed specifically for Stx2a. Although, Stx2a and Stx2c were reported to vary in their affinities to Gb3 in vitro, we expected this assay to detect Stx2c because the primary antibody used targets at the A subunit of Stx2, which is conserved between the 2 subtypes.23,37 Therefore, we next evaluated whether this assay could be applied to measure the Stx2 levels in bacteria lysates from phylogenetically distinct strains. Thirteen strains from 6 of the known nine clades (Table 1) were chosen. These strains consisted of four defined Stx-allelic types: Stx2a only, Stx2c only, Stx1 & 2c and Stx2a & 2c previously.7 One strain (PA7) was previously reported to be Stx1 and Stx2 negative by polymerase chain reaction (PCR), although it was toxin-positive using Premier® EHEC that simultaneously detected both isoforms; and another strain (PA48) could not be categorized by lineage subtype method. We quantified Stx2 in eleven of the thirteen strains (Figure 3), with strains PA7 and PA48 under the detection limit. Therefore, this assay could be applied to detect Stx2c levels in bacteria lysates of strains harboring either Stx2c (PA38, 40) or both Stx2c and Stx1 (PA22, 41).
| Table 1: Characteristics Summary of STEC Strains Used in this Study. |
|
Clade type
|
Toxin profile |
Strain name |
Lineage type*
|
|
Clade 2
|
Stx2a |
PA 32 |
lineage I |
| Clade 2 |
Stx1 & 2a |
PA 5, 31, 49 |
lineage I
|
|
Clade 5
|
Stx1 & 2c |
PA 22 |
lineage I/II |
| Clade 5 |
Stx2c |
PA 38 |
lineage I/II
|
|
Clade 7
|
Stx2c |
PA 40 |
lineage II |
| Clade 7 |
Stx1 & 2c |
PA 41 |
lineage II
|
|
Clade 8
|
Stx1 |
PA 24 |
lineage I/II |
| Clade 8 |
Stx2a & 2c |
PA 35, 39, 47 |
lineage I/II
|
|
Clade 9
|
Stx1 & 2c |
PA 48 |
Undefined |
| Unclassified |
Unknown |
PA7 |
lineage I
|
Figure 3: Quantification of Stx2 Production by E. coli O157:H7 Strains from Various Lineages, Clades, and Toxin Profiles. The Lineage or Clade Type is Shown below the Strain Designation, while the Stx Profile is Differentiated by Color. Data is Reported as the Average Concentration±One Standard Error. Means that do not Share a Common Letter (a, b, c) are Significantly Different (Turkey’s test, p<0.05).
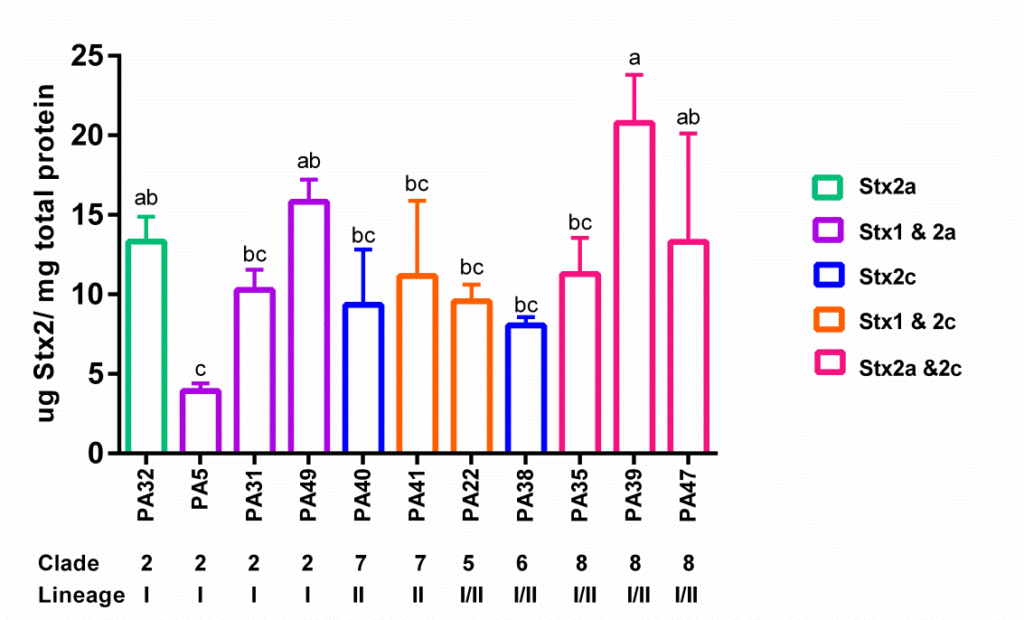
Among them, the highest producer (PA39) had approximately five times more Stx2 production than the lowest strain (PA5). Strains sharing the same lineage, clade or Stx allelic type showed varied Stx2 levels as well. For example, strains PA32, 49 from lineage I produced similar amounts of Stx2 as PA31, but significantly more than lineage I strain PA5. Likewise, among the three strains in clade 8, the Stx2 level in strain PA39 was similar to PA47; however, significantly higher than PA35. Lastly, among strains producing both Stx1 and Stx2a, strain PA49 produced significantly higher amounts of toxin than strain PA5, but was not different from PA31.
DISCUSSION
Stx2a and Stx2c are responsible for severe clinical complications such as life-threatening HUS caused by foodborne pathogen E. coli O157:H7. Thus, the accurate measurement of Stx2 production is important to study its pathogenesis. A globotetraosylceramide (Gb4)-based ELISA was reported for quantitative detection of Stx2e38; however, it can not be applied to measure other Stx2 subtypes because the latters preferentially recognize different receptor, Gb3. Therefore, no quantitative receptor-based ELISA was available for measuring Stx2a before our study by using all commercial available agents. In this study, we have developed a robust assay for the quantification for Stx2a, and it is also expected to detect Stx2c. The cost for this quantitative assay is one third of qualitative Premier® EHEC kit.
Although, Gb3 have been used previously in immunoassays to capture Stx,35,36 we identified several steps that were critical to follow in order to achieve repeatable results. In addition, there are three other modifications from the literature worth highlighting. First, it was reported that the lysoform of Gb3 provided more reliable Stx1 detection, presumably because it is less hydrophobic.36 However, during assay development we found that lyso-CTH was less effective at capturing Stx2a than hydroxyl acyl CTH. When testing the same bacteria lysate from O157:H7 strain with a high Stx2a concentration, the hydroxyl acyl CTH gave saturated reading while the A450 for lyso-CTH is as low as negative control (Table S1). This could simply reflect differences in affinity of Stx1 and Stx2a for CTH derivatives. Secondly, another group previously reported that they could improve R-ELISA sensitivity by increasing the amount of receptor used to coat wells.36 While we saw similar results in initial trials, we decided that 25 µg/ml hydroxyl acyl CTH was the optimal concentration for detecting Stx2a. Under this concentration, we were able to detect Stx2a as low as 23 ng/mL which met our need. In Gb4-based quantitative ELISA for Stx2e, the limit of detection is 20 ng/mL,38 as comparable as ours. Moreover, two 96 well R-ELISA plates can be prepared from one vial of commercial available CTH (0.5 mg), instead of one 96 well plate if a higher concentration was used. Lastly, it was reported that the presence of cholesterol alone or along with lecithin caused statistically significant increases in the binding of Gb3 to Stx2.39 However, no enhancement of Stx2a capture was found when combining these with hydroxyl acyl CTH during the coating step.
Using R-ELISA, we were able to quantify toxin levels from various E. coli O157:H7 strains, and obtained results that were both confirmed and contrasted with those previously reported. First, we did not notice a correlation between Stx2 production and lineage. It was previously reported that E. coli O157:H7 isolates from lineage II produced less Stx2 than those from lineage I and I/II28; however, our findings show that the Stx2 levels in strains from lineage II (PA40, 41) were not significantly different from strains belonging to lineage I (PA5, 31, 32, 49) or lineage I/II (PA22, 35, 38, 47). Second, although strains from clade 8 were suggested to produce higher Stx2 levels than isolates from other clades,29,30 our data indicates this is not universal. In our study, clade 8 strain PA35 produced similar amounts of Stx2 as others clades, including clade 2 (PA31), clade 5 (22), clade 6 (PA38) and clade 7 (PA40, 41). Lastly, it was previously suggested that Stx2-only producing strains synthesized more toxin than Stx1/Stx2-producers, due to crosstalk between phage-encoded repressors encoded in different Shiga toxin-converting phages.40 However, the Stx2a-only strain PA32 produced toxin level that was similar to most Stx1/Stx2-producing strains we tested. It appears likely that phylogeny or toxin profile alone does not explain differences in toxin levels and thus virulence differences observed among strains. More factors such as colonization, adherence capabilities and other virulence genes, need to be considered as well.
Two strains, PA7 and PA48, produced Stx2 levels that were below the detectable limit using our R-ELISA. Strain PA48 is classified as clade 9, and was previously shown to be LSPA type 311111.7 This strain is related to E. coli O157:H7 strain G5101,41 which is also LSPA 31111142 and is atypical among pathogens of this serotype in that it is β-glucuronidase-positive. Little is known about the toxin production levels of these strains. PA7 is a strain previously characterized7 and identified as PCRnegative for both Stx1 and Stx2; however, the commercial kit that simultaneously detects both subtypes identified this strain as Shiga toxin-positive. Our results here suggest the commercial assay was detecting Stx1, although it is possible that PA7 produces Stx2 level below the limit of detection of our R-ELISA.
While this assay is useful for quantifying Stx2a in the bacterial lysates, we have observed limitations when applying it to environmental samples. In a previous publication,43 we were unsuccessful using this method to quantify toxin levels in cattle mucus; however, we could accomplish this using the Premier EHEC kit sold by Meridian Biosciences, Cincinnati, OH, USA. We also found that our assay was one tenth less sensitive than the commercial kit for mouse feces. The sensitivity of the ELISA may be improved by using more hydroxyl acyl CTH as the receptor, or optimizing sample preparation steps to remove inhibitory components from these samples. Therefore, this R-ELISA may not be practical at the moment for detecting toxin in complex biological matrixes. However, it is advantageous when one needs to quantify toxin from laboratory samples while it is impractical to generate or obtain antibodies needed for a traditional sandwich ELISA.
CONCLUSION
Our R-ELISA provides a reliable way to quantify Stx2a using commercially available components, and it can also be used for detecting Stx2c. The application of this assay can be expanded to other Stx2 subtype once corresponding pure toxin is available. It is a readily easy and cheap way to quantify toxin levels especially for groups that currently assess toxin production by qPCR or other transcription-based methods. We predict a similar quantitative assay can be developed to detect Stx1, although as stated above this may require selecting a different receptor mimic.
ACKNOWLEDGEMENTS
We thank Joselyn N. Allen (Pennsylvania State University) for creating the scientific illustration for R-ELISA in this publication; Hillary M. Figler for proofreading this manuscript; Matreya LLC for technical assistance. This work was funded by USDA-NIFA grant 2010-65201-20619.
CONFLICTS OF INTEREST
The authors declare that they have no conflicts of interest.